|
Parameters:Ĭredentials – credentials from get_omero_credentials.īioformats. Use the session ID from an existing login as credentials. Instead of the one included with python-bioformats. To use the matching version of the Bio-formats library for the OMERO release Ice.jar, ice-glacier2.jar, ice-storm.jar and ice-grid.jar. For OMERO server 5.0.0, these are blitz.jar, common.jar, To put the JAR files for your OMERO server version onto your Java classpath Python-bioformats can load images from OMERO URLs. 
PT_BIT = u'bit' ¶Įrrors can be ‘strict’, ‘replace’ or ‘ignore’ and defaults to ‘strict’. PT_UINT8 = u'uint8' ¶Įrrors can be ‘strict’, ‘replace’ or ‘ignore’ and defaults to ‘strict’. 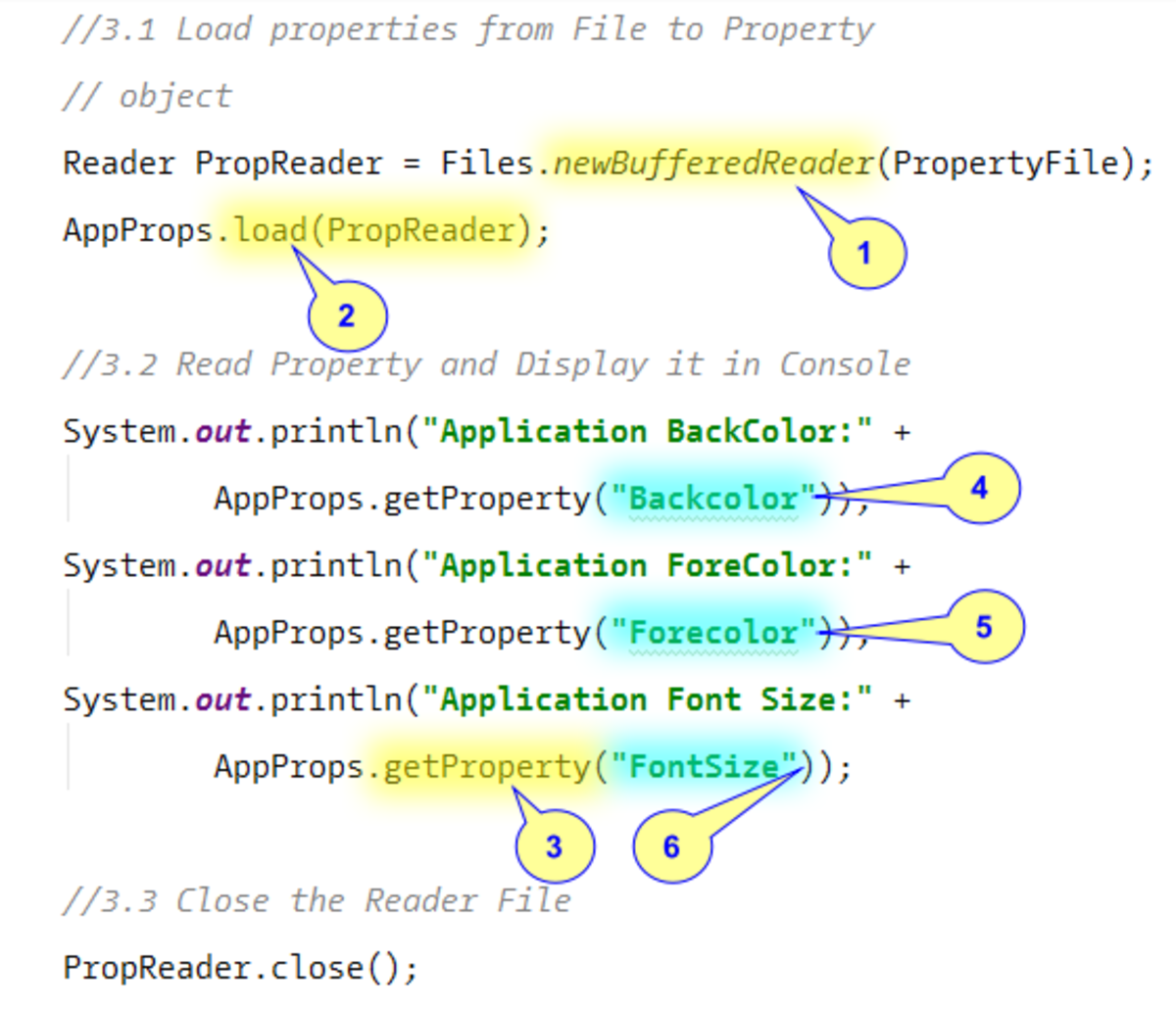
Unicode(string]) -> unicode objectĬreate a new Unicode object from the given encoded string.Įncoding defaults to the current default string encoding.Įrrors can be ‘strict’, ‘replace’ or ‘ignore’ and defaults to ‘strict’. channel_names – names of the channels (make up names if not present).pixel_type – save using this pixel type.write_image ( pathname, pixels, pixel_type, c=0, z=0, t=0, size_c=1, size_z=1, size_t=1, channel_names=None ) ¶ Where it’s supported is to use properties on OMEXML and on some of its Root_node of the DOM and explore it yourself using the DOM API It programatically and get the modified OME-XML back out by calling to_xml. Get a bland OME-XML object which has a one-channel image. Or unicode string, the constructor will parse it and will use it as theīase for any inspection and modification. There are two ways to invoke the constructor. Let the caller create and modify OME-XML. OME-XML, to provide a structured mechanism for inspecting OME-XML and to The OMEXML class has four main purposes: to parse OME-XML, to output Reads and writes OME-XML with methods to get and set it. groupfiles – utilize the groupfiles option to take the directory structureĬlass bioformats.Read the OME metadata from a file using Bio-formats Parameters: get_omexml_metadata ( path=None, url=None ) ¶ load_image_url ( url, c=None, z=0, t=0, series=None, index=None, rescale=True, wants_max_intensity=False, channel_names=None ) ¶ channel_names – None if you don’t want them, a list which will be filled if you do.Įither a 2-d (grayscale) or 3-d (2-d + 3 RGB planes) image.īioformats.t – the frame index in the time dimension.z – the frame index in the z (depth) dimension.Load the given image file using the Bioformats library. load_image ( path, c=None, z=0, t=0, series=None, index=None, rescale=True, wants_max_intensity=False, channel_names=None ) ¶ URL and use it to read an image: bioformats. channel_names – provide the channel names for the OME metadataĬonvenience functions that create an image reader for a file path or.Return a tuple of image and max intensity 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
